






Unless you’ve been living at the bottom of the ocean, you’ll be aware: the world is in the midst of an antibiotic crisis.
This is due to disease-causing bacteria developing resistance to commonly used clinical antibiotics, rendering these antibiotics ineffective. This is known as antimicrobial resistance, or AMR, and is one of the largest global threats that humans face today. It is predicted that by 2050, AMR could result in 10 million deaths globally per year.
Antibiotics (the word itself meaning “anti-life”), are compounds which are capable of killing bacteria and have revolutionized human medicine since their discovery with penicillin in 1928. Suddenly doctors had the ability to prevent and treat bacterial infections which would otherwise have been a death-sentence for the inflicted individual.
We have since becoming hugely reliant on antibiotics to treat everything from simple skin infections to deadly diseases such as tuberculosis, malaria and pneumonia. This is aside from the prophylactic use of antibiotics, common with surgeries such as caesareans. The vast majority of the global population will use antibiotics at least once in their lifetime and many individuals in the US and Europe will take antibiotics multiple times throughout their life. Unfortunately, the overuse of antibiotics is directly responsible for AMR.
.jpg) Bacteria grow and mutate rapidly, and through random mutation can acquire resistance to a particular antibiotic. Use of this antibiotic creates a selective pressure that benefits this bacterium; all the other antibiotic-susceptible bacteria will be killed off, whilst the resistant bacteria will thrive and divide to create more resistant bacteria. The result is an entire population of antibiotic resistant bacteria that are enormously difficult to treat. Even more frightening is that these bacteria can transfer this resistance to other bacteria, something which is known as horizontal gene transfer (HGT).
Bacteria grow and mutate rapidly, and through random mutation can acquire resistance to a particular antibiotic. Use of this antibiotic creates a selective pressure that benefits this bacterium; all the other antibiotic-susceptible bacteria will be killed off, whilst the resistant bacteria will thrive and divide to create more resistant bacteria. The result is an entire population of antibiotic resistant bacteria that are enormously difficult to treat. Even more frightening is that these bacteria can transfer this resistance to other bacteria, something which is known as horizontal gene transfer (HGT).
Bacteria can also acquire resistance to multiple antibiotics, known as multi-drug resistance or MDR. Good examples of this are tuberculosis and MRSA. MRSA stands for methicillin-resistance Staphyloccus aureus, a type of bacteria which can cause severe infection in humans and is often in the news. MRSA is resistant not only to methicillin, but antibiotics oxacillin, amoxicillin and penicillin, making it very challenging to treat. Tuberculosis, caused by the bacteria Mycobacterium tuberculosis, is also increasingly difficult to treat, as strains of Mycobacterium tuberculosis are emerging which are resistant to nearly all antibiotics which can be used to treat it.
As a result, there is enormous effort being put into the discovery of new antibiotics which are effective against these drug-resistant microbes... This brings us to Actinobacteria.
Actinobacteria are bacteria which are found ubiquitously in soil across terrestrial, fresh-aquatic and marine environments where they produce a wide range of antimicrobial compounds to compete with other micro-organisms. The compounds constitute approximately 70% of the world’s antibiotics, as well as many anti-cancer, anti-anthelmintic, anti-viral, anti-hypertensive and anti-fungal compounds. During the 1960-70s, a period known as the ‘Golden Age; of antibiotic discovery, novel antimicrobial compounds were discovered with astonishing frequency from Actinobacteria. In fact, over half the antibiotics in use today were discovered during this period. However, novel antibiotic discovery is now rare, as terrestrial strains of Actinobacteria have been extensively explored and exploited. Re-isolation of the same species and compounds is commonplace and whilst other methods have focused on producing synthetic antibiotics, these have been largely unsuccessful.
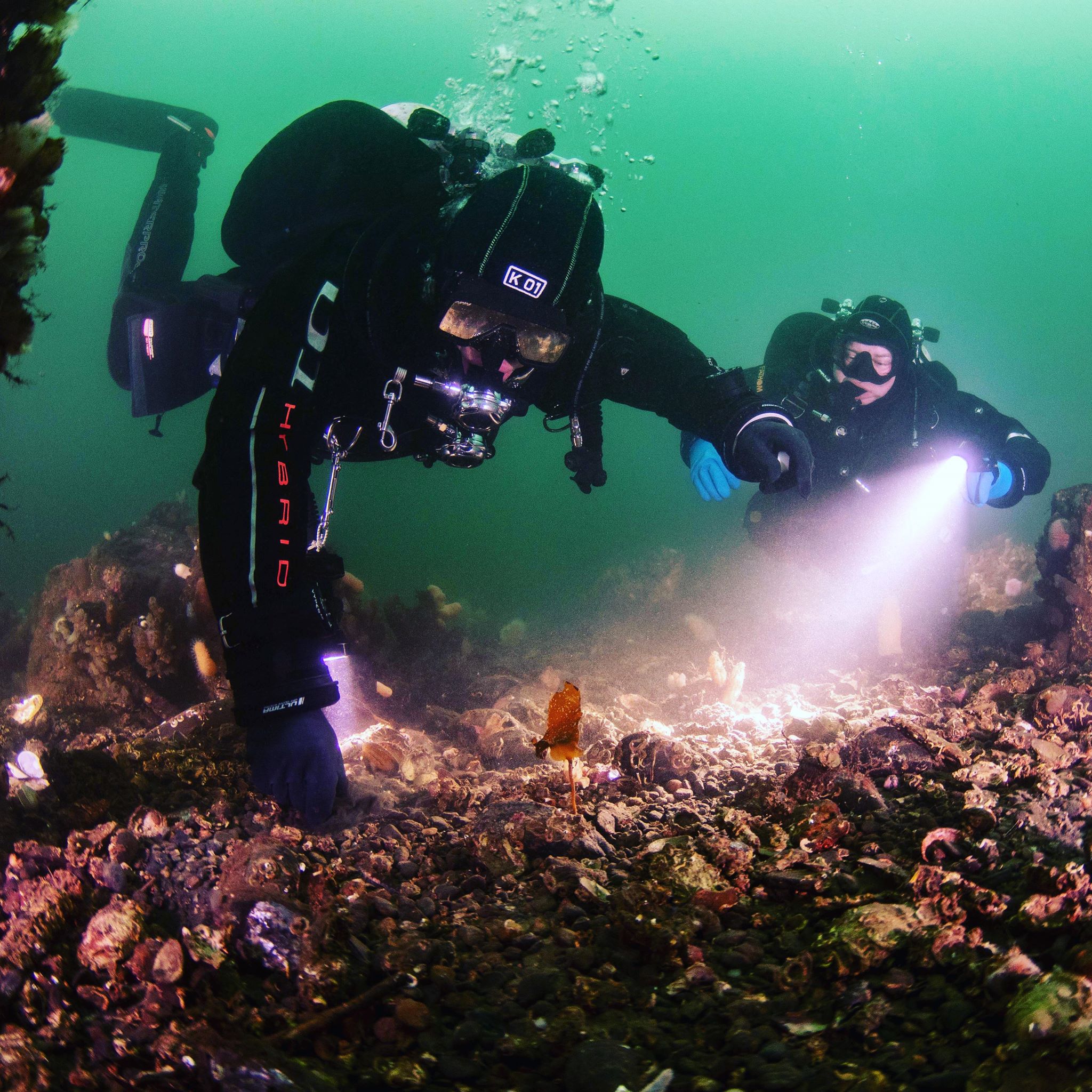

In recent years, the search for new antibiotics has moved to the ocean, which covers 70% of the planet and is the most diverse of the Earth’s environments. Unlike terrestrial sources, the marine biosphere has been relatively under sampled and holds enormous potential for the discovery of new Actinobacteria species which may produce novel antibiotics. One example of this is Salinispora, a type of Actinobacteria which can only survive in marine environments and which was discovered with the help of divers in 1989. Salinispora has proved to be prolific producer of antimicrobials, some of which are effective against MRSA.
It was this that inspired the Dive for Antibiotics (DFA) project, which is funded by the Microbiology Society and began at the Strathclyde Institute of Pharmacy and Biomedical Sciences in 2018. Our project aims are to explore the marine environment for novel Actinobacteria which may produce unknown antimicrobial compounds, whilst also providing education about AMR, Actinobacteria and antibiotic discovery.
It is fairly simple for researchers to collect soil samples from terrestrial environments, but sampling marine environments is far more challenging and requires the help of divers. The DFA project relies on volunteer divers willing to collect sediment samples whilst on their dives and post them to the Hoskisson lab at Strathclyde Institute of Pharmacy and Biomedical Sciences. Participating divers have allowed us to examine marine sediment from all around the UK and from depths as great as 80 meters.
When the project began, we had a few deeply committed divers who would collect samples for us to analyze. However, in our hope to engage more divers with the project, we started on social media on Instagram and Facebook and organized the Dive for Antibiotics competition. Many of our volunteer divers use Halcyon Dive System gear and Halcyon kindly offered their Infinity System as a first-place prize. The project and competition had a fantastic response, with divers clearly being informed individuals who cared about the marine environment, were interested in marine microbes, and excited at the prospect of aiding in antibiotic discovery.
The DFA project sent out a number of sample packs which contained sampling containers and instructions for divers on how to collect their sample. It boiled down to a very simple request for marine sediment – from anywhere in the UK and from any depth. Divers provided the co-ordinates of their sample and the depth taken, information which would allow us to re-examine the area if the sample proved valuable.
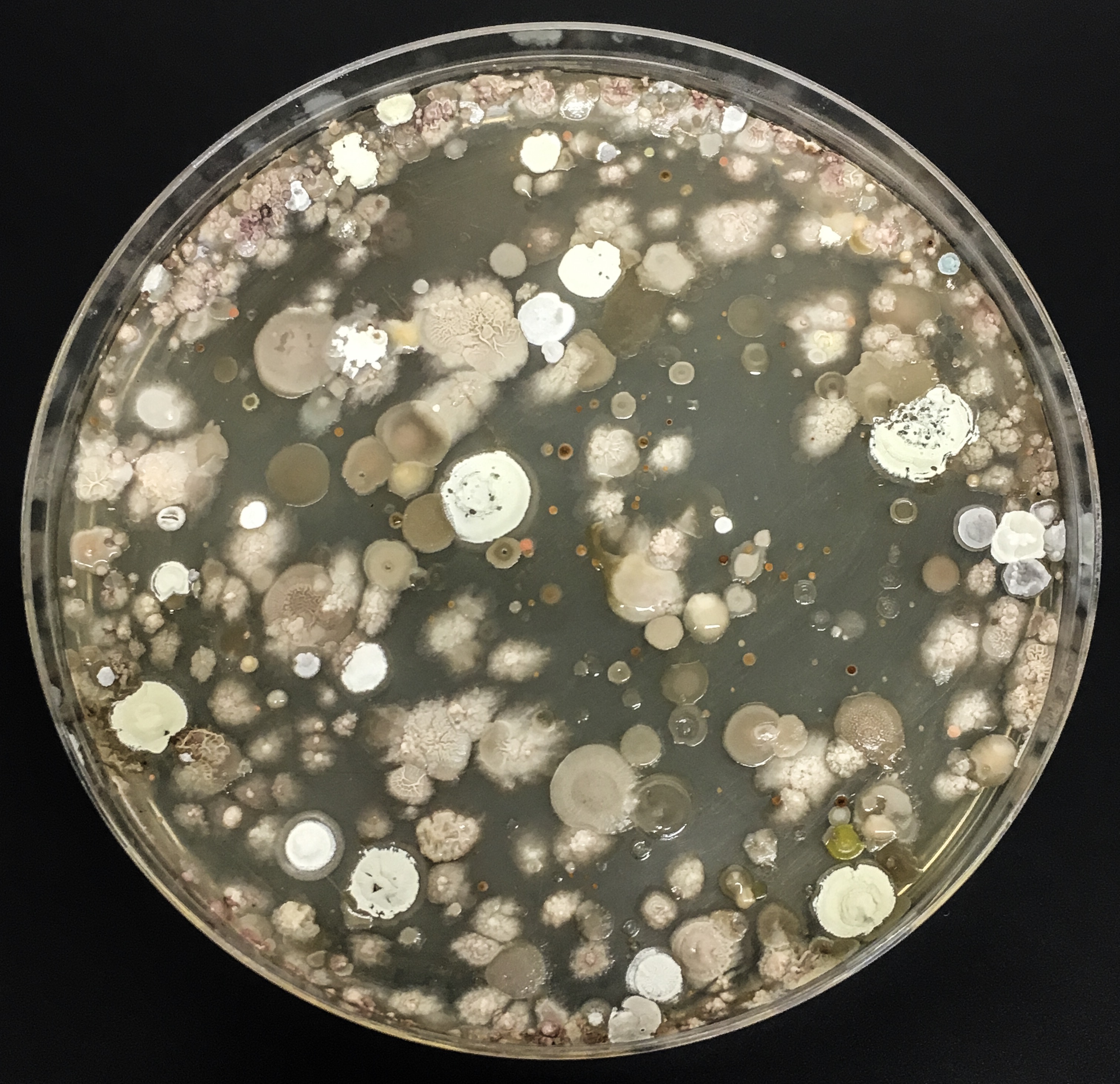 The Actinobacteria isolated are tested for antimicrobial activity; the ability to kill disease causing bacteria such as MRSA. Any Actinobacteria which show antimicrobial activity have their DNA sequenced to determine whether they are a new species of Actinobacteria which may produce novel antimicrobial compounds.
The Actinobacteria isolated are tested for antimicrobial activity; the ability to kill disease causing bacteria such as MRSA. Any Actinobacteria which show antimicrobial activity have their DNA sequenced to determine whether they are a new species of Actinobacteria which may produce novel antimicrobial compounds.
The samples returned to our lab came from areas all around Scotland, England and Ireland and produced numerous interesting Actinobacteria isolates. Many divers collected their samples from well-known shipwrecks, such as those at Scapa Flow in the Orkney Islands. However, one of our most interesting isolates was derived from a scallop bed off the Isle of Skye, Scotland.
So far, testing has shown that 5 Actinobacteria isolates collected as part of the DFA competition are showing anti-microbial activity. One in particular shows bactericidal activity against P. aeruginosa, which is particular difficult to kill. Others are producing compounds which are capable of killing MRSA.
Our research is currently on going, and we hope to identify the species of our isolates and determine whether the anti-microbials they are producing are novel in nature. The DFA project also hopes to continue engaging passionate divers in our research and spreading information about antibiotic discovery and the importance of the marine microbial environment.
For more information, follow this link: https://www.instagram.com/diveforantibiotics/?hl=en
|
Halcyon Dive Systems 1.800.HALCYON (425.2966) 1 (386) 454-0811 |
|
Halcyon Dive Systems 24587 NW 178th Place High Springs, FL 32643 |
|
Customer Service Hours Monday-Thursday 8:00AM - 4:30PM EST Friday 8:00AM - 11:00AM EST |